Challenge Open Access
AF Classification from a Short Single Lead ECG Recording: The PhysioNet/Computing in Cardiology Challenge 2017
Gari D. Clifford , Chengyu Liu , Benjamin Moody , Li-wei Lehman , Ikaro Silva , Alistair Johnson , Roger Mark
Published: Feb. 1, 2017. Version: 1.0.0
Community forum for the 2017 PhysioNet/CinC Challenge (April 28, 2018, 2:10 p.m.)
If you have any questions or comments regarding this challenge, please post it directly in our Community Discussion Forum. This will increase transparency (benefiting all the competitors) and ensure that all the challenge organizers see your question.
2017 PhysioNet Challenge Papers Available (Feb. 2, 2018, midnight)
All the papers from the 2017 Challenge along with their corresponding PDFs are now available.
More news
Slides describing the results of the 2017 PhysioNet/CinC Challenge (Oct. 23, 2017, 2:09 p.m.)
Slides presented at Computing in Cardiology in Rennes, France on the 26th-27th September 2017, and at the Annual Data Institute Conference at the USF Data Science Institute, San Francisco, CA, USA on the 17th October 2017 can be found here .
Performance of algorithms in the 2017 PhysioNet/CinC Challenge (Oct. 10, 2017, 2:06 p.m.)
A more detailed breakdown of performance of each official algorithm is now available here.
Official results for the 2017 PhysioNet/CinC Challenge (Oct. 10, 2017, 2 p.m.)
The official results are now available, as well as a summary paper of the Challenge.
Winners announced for the 2017 PhysioNet/CinC Challenge (Sept. 27, 2017, 1:57 p.m.)
Winners were announced at Computing in Cardiology 2017 in Rennes, France! Four teams were tied for first place, with an overall score of 0.83:
- Shreyasi Datta, Chetanya Puri, Ayan Mukherjee, Rohan Banerjee, Anirban Dutta Choudhury, Arijit Ukil, Soma Bandyopadhyay, Rituraj Singh, Arpan Pal, and Sundeep Khandelwal
- Shenda Hong, Meng Wu, Yuxi Zhou, Qingyun Wang, Junyuan Shang, Hongyan Li, and Junqing Xie
- Tomás Teijeiro, Constantino A. García, Paulo Félix, and Daniel Castro
- Morteza Zabihi, Ali Bahrami Rad, Aggelos K. Katsaggelos, Serkan Kiranyaz, Susanna Narkilahti, and Moncef Gabbouj
PhysioNet/CinC Challenge 2017 (Feb. 1, 2017, 5 p.m.)
We are excited to announce the opening of the annual PhysioNet/Computing in Cardiology Challenge for 2017: AF Classification from a short single lead ECG recording. A database of over 10,000 ECG recordings is being made freely available exclusively for this competition by AliveCor, and represent a serendipitous sample of patient-initiated recordings of one minute or less. They have all been labeled for rhythm by hand into one of four categories: Normal, Atrial Fibrillation, Other Rhythm or Too Noisy to Process. We challenge the public to develop the most accurate classifier of these data into these four categories. As usual, a portion of these data have been hidden from the public to allow us to objectively assess your algorithms.
Please include the standard citation for PhysioNet:
(show more options)
Goldberger, A., Amaral, L., Glass, L., Hausdorff, J., Ivanov, P. C., Mark, R., ... & Stanley, H. E. (2000). PhysioBank, PhysioToolkit, and PhysioNet: Components of a new research resource for complex physiologic signals. Circulation [Online]. 101 (23), pp. e215–e220.
Introduction
The 2017 PhysioNet/CinC Challenge aims to encourage the development of algorithms to classify, from a single short ECG lead recording (between 30 s and 60 s in length), whether the recording shows normal sinus rhythm, atrial fibrillation (AF), an alternative rhythm, or is too noisy to be classified.
There are various types of cardiac arrhythmias that may be classified by:
- Origin: atrial arrhythmia, junctional arrhythmia, or ventricular arrhythmia.
- Rate: tachycardia ( > 100 beats per minute (bpm) in adults) or bradycardia ( < 60 bpm in adults).
- Mechanism: automaticity, re-entry, triggered.
- AV Conduction: normal, delayed, blocked.
- Duration: non-sustained (less than 30 s) or sustained (30 s or longer).
AF is defined as a “tachyarrhythmia characterized by predominantly uncoordinated atrial activation with consequent deterioration of atrial mechanical function” by the American College of Cardiology (ACC), the American Heart Association (AHA) and the European Society of Cardiology (ESC) [1]. AF is the most common sustained cardiac arrhythmia, occurring in 1-2% of the general population [2; 3] and is associated with significant mortality and morbidity through association of risk of death, stroke, hospitalization, heart failure and coronary artery disease, etc. [3; 4]. More than 12 million Europeans and North Americans are estimated to suffer from AF, and its prevalence will likely triple in the next 30-50 years [5]. More importantly, the incidence of AF increases with age, from less than 0.5% at 40-50 years of age, to 5-15% for 80 year olds [6].
Despite the enormity of this problem, AF detection remains problematic, because it may be episodic. AF detectors can be thought of belonging to one of two categories: atrial activity analysis-based or ventricular response analysis-based methods. Atrial activity analysis-based AF detectors are based on the analysis of the absence of P waves or the presence of fibrillatory f waves in the TQ interval. Published methods to do this include: an echo state neural network [7], P-wave absence (PWA) based detection [8], analysis of the average number of f waves [9], P-wave-based insertable cardiac monitor application [10], wavelet entropy [11] [12] and wavelet energy [13]. Atrial activity analysis-based AF detectors can achieve high accuracy if the recorded ECG signals have little noise contamination and high resolution, but tend to suffer disproportionately from noise contamination [4]. In contrast, ventricular response analysis is based on the predictability of the inter-beat timing (‘RR intervals’) of the QRS complexes in the ECG. RR intervals are derived from the most obvious large amplitude feature in the ECG, the R-peak, the detection of which can be far more noise resistant. This approach may therefore be more suitable for automatic, real-time AF detection [14]. Published methods include: Poincaré plot analysis [15], Lorenz plot analysis [16], analysis of cumulative distribution functions [17], thresholding on the median absolute deviation (MAD) of RR intervals [18], histogram of the first difference of RR intervals [19], minimum of the corrected conditional entropy of RR interval sequence [20], 8-beat sliding window RR interval irregularity detector [21], symbolic dynamics and Shannon entropy [22], sample entropy of RR intervals [23; 24; 25], and normalized fuzzy entropy of RR intervals [26].
It is worth noting that AF detectors that combine both atrial activity and ventricular response could provide an enhanced performance by combining independent data from each part of the cardiac cycle. Such detection approaches have included: RR interval Markov modeling combined with PR interval variability and a P wave morphology similarity measure [27] and a fuzzy logic classification method which uses the combination of RR interval irregularity, P-wave absence, f-wave presence, and noise level [28]. It is also worth noting that multivariate approaches based on machine learning that combines several of the above single features can also provide enhanced AF detection [29; 30; 31].
Previous studies concerning AF classification are generally limited in applicability because 1) only classification of normal and AF rhythms were performed, 2) good performance was shown on carefully-selected often clean data, 3) a separate out of sample test dataset was not used, or 4) only a small number of patients were used. It is challenging to reliably detect AF from a single short lead of ECG, and the broad taxonomy of rhythms makes this particularly difficult. In particular, many non-AF rhythms exhibit irregular RR intervals that may be similar to AF. In this Challenge, we treat all non-AF abnormal rhythms as a single class and require the Challenge entrant to classify the rhythms as 1) Normal sinus rhythm, 2) AF, 3) Other rhythm, or 4) Too noisy to classify.
Quick Start
- Download the training set:
training2017.zipand the sample MATLAB entry:sample2017.zip. - Develop your entry by making the following edits to
sample2017.zip:- Modify the sample entry source code file
challenge.mwith your changes and improvements. - Modify the
AUTHORS.txtfile to include the names of all the team members. - Unzip
training2017.zipand move all its files to the top directory of your entry directory (wherechallenge.mis located). - Run your modified source code file on the validation records in the training set by executing the script
generateValidationSet.m. This will also build a file calledentry.zip. - Optional: Include a file named
DRYRUNin the top directory of your entry (where theAUTHORS.txtfile is located) if you do not wish your entry to be scored and counted against your limit. This is useful in cases where you wish to make sure that the changes made do not result in any error.
- Modify the sample entry source code file
- Submit your
entry.zipfor scoring through the PhysioNet/CinC Challenge 2017 project (Update: submissions are now closed). The contents ofentry.zipmust be laid out exactly as in the sample entry. Improperly-formatted entries will not be scored.
Join our community Community Discussion Forum to get the latest challenge news, technical help, or if you would like to find partners to collaborate with.
Rules and Deadlines
Participants are asked to classify one lead ECG recordings as normal rhythm, AF, other rhythm or noisy recordings.
Entrants may have an overall total of up to 15 submitted entries over both the unofficial and official phases of the competition (see Table 1). Each participant may receive scores for up to five entries submitted during the unofficial phase and ten entries at the end of the official phase. Unused entries may not be carried over to later phases. Entries that cannot be scored (because of missing components, improper formatting, or excessive run time) are not counted against the entry limits.
All deadlines occur at noon GMT (UTC) on the dates mentioned below. If you do not know the difference between GMT and your local time, find out what it is before the deadline!
| Start at noon GMT on | Entry limit | End at noon GMT on | |
|---|---|---|---|
| Unofficial Phase | 1 February | 5 | 9 April |
| [Hiatus] | 9 April | 0 | 16 April |
| Official Phase | 16 April | 10 | 1 September |
All official entries must be received no later than noon GMT on Friday, 1 September 2017. In the interest of fairness to all participants, late entries will not be accepted or scored. Entries that cannot be scored (because of missing components, improper formatting, or excessive run time) are not counted against the entry limits.
To be eligible for the open-source award, you must do all of the following:
- Submit at least one open-source entry that can be scored before the Phase I deadline (noon GMT on Sunday, 9 April 2017).
- Submit at least one entry during the second phase (between noon GMT on Sunday, 16 April 2017 and noon GMT on Friday, 1 September 2017). Only your final entry will count for ranking.
- Entering an Abstract to CinC: Submit an acceptable abstract (about 299 words) on your work on the Challenge to Computing in Cardiology no later than 15 April 2017. Include the overall score for at least one Phase I entry in your abstract. Please select “PhysioNet/CinC Challenge” as the topic of your abstract, so it can be identified easily by the abstract review committee. You will be notified if your abstract has been accepted by email from CinC during the first week in June.
- Submit a full (4-page) paper on your work on the Challenge to CinC no later than the deadline of conference paper submission.
- Attend CinC 2017 (24–27 September 2017) in Rennes, France and present your work there.
Please do not submit analysis of this year’s Challenge data to other Conferences or Journals until after CinC 2017 has taken place, so the competitors are able to discuss the results in a single forum. We expect a special issue from the journal Physiological Measurement to follow the conference and encourage all entrants (and those who missed the opportunity to compete or attend CinC 2017) to submit extended analysis and articles to that issue, taking into account the publications and discussions at CinC 2017.
Challenge Data
ECG recordings, collected using the AliveCor device, were generously donated for this Challenge by AliveCor. The training set contains 8,528 single lead ECG recordings lasting from 9 s to just over 60 s (see Table 2) and the test set contains 3,658 ECG recordings of similar lengths. The test set is unavailable to the public and will remain private for the purpose of scoring for the duration of the Challenge and for some period afterwards.
ECG recordings were sampled as 300 Hz and they have been band pass filtered by the AliveCor device. All data are provided in MATLAB V4 WFDB-compliant format (each including a .mat file containing the ECG and a .hea file containing the waveform information). More details of the training set can be seen in Table 2. Figure 1 shows the examples of the ECG waveforms (lasting for 20 s) for the four classes in this Challenge. From top to bottom, they are ECG waveforms of normal rhythm, AF rhythm, other rhythm and noisy recordings.
Please note, since all the classification was performed by a single expert, we are currently in the process of re-scoring a subset where our trust of the data is lowest. We will almost certainly update labels on some of data in both the training and test databases. We may also add new data in the future, although this is unlikely to change after we begin the official phase of the competition.
Please also note that the scoring system currently treats all classes equally. It is likely we will update this for the official phase also. The point of the unnofficial phase is for us to iron out bugs in the competition, and you are a vital part of that. Please send us suggestions/queries to challenge@physionet.org. We welcome suggestions for reclassifying a file, but please provide your reasoning.
We strongly suggest you use the Google group public forum to ask questions unless you are asking us a question that is specific to your entry only and would reveal your methods to others. In general we post answers to the Google group to provide a level playing field.
Please note that we are all volunteers with finite bandwidth, so we prioritize our responses and often discuss them as a group first. Inevitably some emails will have a response delay or may even slip through the net.
| Type | # recording | Time length (s) | ||||
|---|---|---|---|---|---|---|
| Mean | SD | Max | Median | Min | ||
| Normal | 5154 | 31.9 | 10.0 | 61.0 | 30 | 9.0 |
| AF | 771 | 31.6 | 12.5 | 60 | 30 | 10.0 |
| Other rhythm | 2557 | 34.1 | 11.8 | 60.9 | 30 | 9.1 |
| Noisy | 46 | 27.1 | 9.0 | 60 | 30 | 10.2 |
| Total | 8528 | 32.5 | 10.9 | 61.0 | 30 | 9.0 |
Figure 1. Examples of the ECG waveforms.
Sample Submission
As a starting point we have provided an example entry (post here: sample2017.zip) which provides a state of the art detector based upon the method described by Sarkar et al [16]. We note that this detector provides a classification of only Normal or AF rhythms. We leave it to the Challengers to add the other classes.
You may want to begin with the sample detector, or discard it completely and start from scratch using more data-driven or physiological model-based approaches. Please note that the sample entry has been patented by their original authors and, although we provide an open source version of it for benchmarking, you should not hope to create intellectual property from derivatives of it. We therefore suggest you concentrate on developing alternative methods to this benchmark.
NOTE: You do not need any additional software, apart from Matlab or GNU Octave, to run the sample entry. You can use any programming languages or libraries you like when implementing your own submission.
Preparing an entry for the challenge
To participate in the challenge, you will need to create software that is able to read the test data and output the final classification result without user interaction in our test environment. One sample entry (sample2017.zip, written in MATLAB) is available to help you get started. In addition to MATLAB, you may use any programming language (or combination of languages) supported using open source compilers or interpreters on GNU/Linux, including C, C++, Fortran, Haskell, Java, Octave, Perl, Python, and R.
If your entry requires software that is not installed in our sandbox environment, we will work with you during Phase I to try to ensure your code can run. We will not modify the test environment after the start of Phase II of the challenge.
Participants should download the sample entry (sample2017.zip). Entries should have the exact layout of the sample entry; specifically, they must contain:
setup.sh, a bash script runs once before any other code from the entry; use this to compile your code as needed.next.sh, a bash script runs once per training or test record; it should analyze the record using your code, saving the results as a text file for each record.dependencies.txt, a text file that lists additional Debian packages that must be installed prior to running your entry’ssetup.shandnext.shscripts.answers.txt, a text file containing the results of running your program on each record in the validation set (part of training set, see below for details). These results are used for validation only, not for ranking entries.AUTHORS.txt, a plain text file listing the members of your team who contributed to your code, and their affiliations.LICENSE.txt, a text file containing the license for your software (the default is the GPL). All entries are assumed to be open source and will eventually be released on PhysioNet (for closed source entries please see below).
See the comments in the sample entry’s setup.sh and next.sh if you wish to learn how to customize these scripts for your entry.
We verify that your code is working as you intended, by running it on a small subset (validation set, 300 recordings) of the training set, then comparing the answers.txt file that you submit with your entry with answers produced by your code running in our test environment using the same records. Using a small portion of the training set means you will know whether your code passed or failed to run within a small time. If your code passes this validation test, it is then evaluated and scored using the hidden test set. The score in the hidden test set determines the ranking of the entries and the final outcome of the Challenge. Note that in the Official Phase of the challenge, more data may be added to both training and hidden test set. Your final entry in the Official Phase of the competition will be run on the entire test set, and so may take much longer than earlier entries.
In addition to the required components, your entry may include a file named DRYRUN. If this file is present, your entry is not evaluated using the hidden test data, and it will not be counted against your limit of entries per phase; you will receive either a confirmation of success or a diagnostic report, but no scores. Use this feature to verify that none of the required components are missing, that your setup.sh script works in the test environment, and that your next.sh script produces the expected output for the training data within the time limits.
Closed Source Entries
Although the competition is only for open source entries, we also accept the submission of closed-source entries from industry or from individuals. If you enter closed source, we will not publish your code or score (unless you specifically request that we do so). However, the default entry is open source (GPL), so you must explicitly indicate that your entry is closed source by including with your entry a file called CLOSEDSOURCE.txt and modifying LICENSE.txt accordingly. If you submit an executable, it must be compiled to run in our testing environment (Debian 8.7, amd64.)
Open source entry scores will not be posted until after the close of the Official Phase, and closed source entries will not be posted. You may choose to swap between being open source or closed source at any time up to the end of the Unofficial Phase by inserting or removing the CLOSEDSOURCE.txt file with your final entry prior to the end of the Unofficial Phase.
More on Licences and IP
We would like to note that the competition does not give the company donating the data any rights to algorithms or ideas developed by competitors. Any entity is free to contact a competitor to request a license to use their code for commercial purposes. Since the competitor must use an open source license to be eligible for a prize, it may be necessary for the competitor to produce another version of the code with a different license (as copyright holders they are at liberty to do so). This would in no way influence the posting of the open source code for the Challenge or its use for research purposes.
Scoring
If your entry is properly formatted, and nothing is missing, it is tested and scored automatically, and you will receive your scores when the test is complete (depending on your entry’s run time, this may take an hour or more). If you receive an error message instead, read it carefully and correct the problem(s) before resubmitting. Missing answers are treated as noise labels.
The scoring for this challenge uses a F1 measure, which is an average of the four F1 values from each classification type. The counting rules for the numbers of the variables are defined in Table 3 as below:
Table 3. Counting rules for the numbers of the variables.
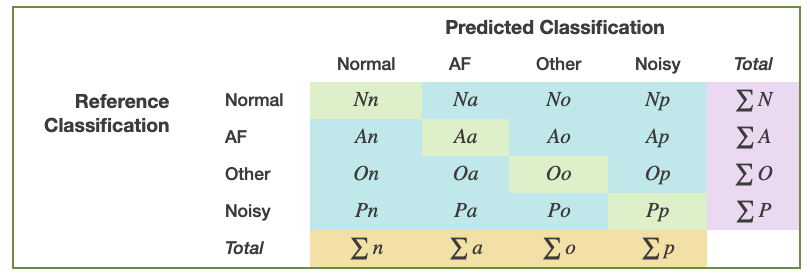
For each of the four types, F1 is defined as:
- Normal rhythm:
- AF rhythm:
- Other rhythm:
- Noisy:
The script score2017Challenge.m in the sample entry, sample2017.zip, generates the final challenge score as follows:
The score on the hidden data represents your algorithm’s performance on a subset of the data. We will assess your performance on all of the hidden data only on your final entry in the official phase of the competition. The reason we do this is to prevent you over training on the hidden data. To improve your algorithm, you must assess your algorithm’s performance on the training data using cross validation.
We are arbitrarily following the olympic model (for swimming events). Anyone with exactly the same score would split the 'medals' as normal. If two tie for gold (1st place) then we skip silver and the prize money for first and second place are divided equally between the 1st place winners. The 3rd place is awarded as normal. If three or more tie for gold then we award no other places and the entire cash prize for all three places is split equally. If any number tie for silver we skip bronze and split the silver and bronze cash prizes equally between all the 2nd place teams. If several tie for bronze then they all are awarded 3rd place share the 3rd place prize money equally.
So if a team scored 0.95 and then two teams scored 0.91, and others scored 0.85 (say) the first prize would go to the team with 0.95 and the second place would be equally awarded to the two teams scoring 0.91. No 'third place' team would be identified.
Note that the third decimal place in the scoring system probably isn't statistically significant, which is why our leader board doesn't show it beyond two decimal places (like they do in olympic swimming events).
Abstract Submission
Don't forget you must also submit an abstract to Computing in Cardiology before the imminent deadline on the 15th of April and attend the conference in September where we will announce the winner. See cinc.org. Note that your methods and score, and therefore your abstract, will most likely change by the end of summer. That is acceptable and expected. If you don't submit now though, you won't reserve your place to discuss your methods at the conference. Please note that abstracts should include your current methods and score. We encourage you to include cross validation stats on the training data too to show the reviewers you know what you are doing. In many ways toys is far more important than your test set score we give you at this stage, so if you have a poor challenge score but a great cross validated score then you are well on your way!
Incoherent or information poor abstracts are unlikely to be accepted as they indicate low quality approaches and an inability to communicate ideas. A well thought out abstract indicates a high likelihood of a good presentation and high quality scientific approach. Do not include a description of the competition in the abstract (it's very clear what the competition is about) ... focus on your methods and results.
Please make sure you select the Physionet Challenge category in the abstract submission. If you don't it may get reviewed outside of the challenge track and get rejected.
Also, please do not title your paper something similar to the challenge itself .... I'll ask you to change your title and it wastes both of our time. To be clear, our paper will be called:
AF classification from a short single lead ECG recording: The Physionet Computing in Cardiology Challenge 2017.
... so please do not include the phrases Physionet, Computing in Cardiology, or Challenge 2017 in your title. Please make your title specific to your technique. E.g.: "AF detection using cepstral coefficients, higher order moments and an SVM" would be fine .... but "deep learning for AF detection" is too general and will make your title indistinguishable from half the entries. Think about what makes your approach different.
Final Entry
All competitors are required to identify which of their entries they believe is their best performing entry by September 6 at 11am UTC. We will then assess this code on the entire hidden data. You will be able to change your selection at any time before September 6. If you do not designate a final entry by September 6 at 11:00 AM, we will use your last successfully-submitted entry.
Papers, Presentations and Posters for CinC
Creating your conference paper
You should post a 99% complete version of your paper at least one week before the conference and mark it openly available for others to read. You obviously can’t insert your final score in there until after the conference, and so you will have 3–4 days after the conference to update your paper. However, you must submit an acceptable and ostensibly complete 4 page paper that will not undergo any revisions after the conference, except to add the final score. Why? Well we don’t want you spending time addressing other groups entries. You don’t have the time to do it, nor will you have enough detailed information to do so. The time to do that is in the follow-on Physiological Measurement special issue, where we expect you to submit a full paper (as long as you like, as long as it’s all pertinent) where you can revisit your algorithm in light of the feedback at the conference, or even improve it by using other competitor’s ideas! It would be unfair to do so in the conference article. However, this is exactly how science progresses (after the conference) , and indeed, we hope that this conference facilitates you to swap ideas, question each other, improve, collaborate, compete if you wish (in a kind spirited manner), publish and share.
We strongly advise you to click the box requesting that your article be pushed to the preprint site for the conference—it will help others understand more clearly what you did, and why you did it (10 minutes or a poster is rarely long enough to explain yourself). Please dedicate your article almost entirely to your methods, including the preprocessing, feature extraction, model pruning, data relabelling, external data sets used, and training cross validation. There is absolutely no need to describe the competition—we will do that in detail and you can directly reference the challenge website and the article we will publish:
Gari Clifford, Chengyu Liu, Benjamin Moody, Li-wei H. Lehman, Ikaro Silva, Qiao Li, Alistair Johnson, Roger G. Mark. AF Classification from a Short Single Lead ECG Recording: the PhysioNet Computing in Cardiology Challenge 2017. Computing in Cardiology (Rennes: IEEE), Vol 44, 2017 (In Press).
We strongly encourage you to reference this article, because it will provide the reader of your article with details missing from the website. We will post a pre-print of this in mid September (at least a week before the conference and possibly two). It will not include the list of winners and the final rankings, although we will add this in the final version immediately after the conference.
Since you do not know who the winner will be, and what your final ranking will be, please do not indicate your position in the rankings until after the competition when the final scores are known. It is misleading to include your rankings either in the unofficial phase or at any time in the official phase. Only the final scores are meaningful, as they will have been assessed on the most accurate version of the (entire) hidden test data.
When creating your 4 page conference article please use this latex example as a template, or download the template from http://cinc.mit.edu/authors_kit/papers/ which also includes a Word template (if you feel the need to use that). Further instructions can be found at that URL, and include details such as capitalization of subheadings, etc. Do not play with the margins, font or other parameters. The PDF generated from the source will be auto-checked for format conformity and rejected if the margins are wrong, it is too long or otherwise contravenes the instructions.
Creating your conference presentation
Be concise—no-one should discuss the aims of the competition or the databases. The challenge organizers will have done that in the first presentation of each session. Instead, focus on what you did—how you trained your algorithm, what features you selected, which open source algorithms you used, and how you stratified the data. Make sure you present cross fold validation results from the training data (as well as your overall ranking in the official competition). Stratify by patient and by database (if you have used external databases). Do not include the scores from the unofficial phase; these are unimportant. Be sure to indicate if your algorithm is open source or not.
Do compare your results to others and be realistic about your performance on the test data. Explain what was novel in your algorithm and what made it work, and also what didn’t work too well.
Use pictures on each slide to draw the eye; please don’t list a series of bullet points and repeat them verbatim. They should be short sentences—just enough to remind you what you wanted to say. The fewer words the better.
Diagrams are beautiful, so do include them ... but please make sure we can read all the captions from the back of the room. Use high contrast thick lines; it’s usually a long room. Try it out in your biggest auditorium before coming to the conference.
Above all, don’t run over; you will look unprofessional and fail to convey your message. Keep it down to 7 slides at most—you only have 10 minutes and you will be cut off after that. (It’s unfair to use up other people’s time, and you must allow a few minutes for questions). You can add slides afterwards to answer questions, or add a postscript if you want.
So—make sure you practice your presentation content and timing before you come! Ask your colleagues to listen to you and critique the slides—if they don’t understand you, we probably won’t.
Creating your poster presentation
If you were selected for a poster presentation, don’t worry—you still have to write a paper which is listed with IEEE Explore and Google Scholar, and it counts as much as if you had presented in an oral session. (There’s no way to retrospectively find out if your 4 page paper was presented orally or as a poster).
The maximum size was 85 cm (33") wide by 115 cm (45") tall for CinC 2016. Note that poster dimensions vary from year to year so don’t assume that a size that fit last year will fit this year. You will receive email instructions from the conference organizers explaining this year’s restrictions. Please make every attempt to build a visually appealing presentation, which conveys the right points.
More information can be found here: http://cinc.mit.edu/authors_kit/presentations/. You may want to read these resources from researchers at Cornell and Oxford.
There is a prize for the best poster, so do take it seriously, and make sure you stand by it during your session if you want to be in the running!
After the Challenge
As is customary, we hope to run a special issue in Physiological Measurement with a closing date of 31 January 2018. We will therefore encourage competitors (and non-competitors) to submit updates and further reworks based on the Challenge after the award ceremony at the Computing in Cardiology Conference in Rennes in September.
Obtaining complimentary MATLAB licenses
The MathWorks has kindly decided to sponsor Physionet’s 2017 Challenge providing both prize money and licenses. The MathWorks is offering to all teams that wish to use MATLAB, complimentary licenses. User can apply for a license and learn more about MATLAB support through The Mathwork’s PhysioNet Challenge link. If you have questions or need technical support, please contact The MathWorks at academicsupport@mathworks.com.
Challenge Results
The top-scoring programs submitted in the PhysioNet/Computing in Cardiology Challenge 2017 are listed here. For more information about the details of these algorithms, see the corresponding papers.
2017 Challenge Entries
* These entries were disqualified because they included code that was not freely licensed, contrary to the Challenge rules. The code in question has been removed from the archives linked above.
† These entries were disqualified because their authors submitted more than ten entries in total, contrary to the Challenge rules.
Papers
The following paper is an introduction to the challenge topic, with a summary of the challenge results and a discussion of their implications. Please cite this publication when referencing the Challenge.
Gari Clifford, Chengyu Liu, Benjamin Moody, Li-Wei Lehman, Ikaro Silva, Qiao Li, Alistair Johnson, Roger Mark. AF Classification from a Short Single Lead ECG Recording: the Physionet Computing in Cardiology Challenge 2017.
Over 40 papers were presented by participants at Computers in Cardiology 2017. These papers have been made available by their authors under the terms of the Creative Commons Attribution License 3.0 (CCAL). See this page for a list of the papers. We wish to thank all of the authors for their contributions.
References
Access
Access Policy:
Anyone can access the files, as long as they conform to the terms of the specified license.
License (for files):
Open Data Commons Attribution License v1.0
Discovery
Topics:
atrial fibrillation
challenge
ecg
Corresponding Author
Files
Total uncompressed size: 0 B.
Access the files
- Download the ZIP file (1.4 GB)
- Access the files using the Google Cloud Storage Browser here. Login with a Google account is required.
-
Access the data using the Google Cloud command line tools (please refer to the gsutil
documentation for guidance):
gsutil -m -u YOUR_PROJECT_ID cp -r gs://challenge-2017-1.0.0.physionet.org DESTINATION
-
Download the files using your terminal:
wget -r -N -c -np https://physionet.org/files/challenge-2017/1.0.0/
-
Download the files using AWS command line tools:
aws s3 sync s3://physionet-open/challenge-2017/1.0.0/ DESTINATION
| Name | Size | Modified |
|---|---|---|
| papers | ||
| sources | ||
| training | ||
| validation | ||
| Clifford_Physionet_Challenge2017_USF_DS_Conference_October_15-17-2017.pdf (download) | 4.0 MB | 2019-04-17 |
| Clifford_et-al-challenge_2017_CinC_paper.pdf (download) | 547.9 KB | 2019-04-17 |
| REFERENCE-v0.csv (download) | 75.0 KB | 2019-04-17 |
| REFERENCE-v1.csv (download) | 75.0 KB | 2019-04-17 |
| REFERENCE-v2.csv (download) | 75.0 KB | 2019-04-17 |
| REFERENCE-v3.csv (download) | 75.0 KB | 2019-04-17 |
| example_waveforms.svg (download) | 93.1 KB | 2019-04-17 |
| followup-combined-sorted-with-ranking-median-95thpc.csv (download) | 20.9 KB | 2019-04-17 |
| followup-combined-sorted-with-ranking-median-95thpc.xls (download) | 112.5 KB | 2019-04-17 |
| python-dummy-entry.zip (download) | 15.4 KB | 2019-04-17 |
| results.csv (download) | 5.5 KB | 2019-04-17 |
| results_all_F1_scores_for_each_classification_type.csv (download) | 8.9 KB | 2019-04-17 |
| sample2017.zip (download) | 3.4 MB | 2019-04-17 |
| table3.png (download) | 40.1 KB | 2020-01-22 |
| training2017.zip (download) | 94.6 MB | 2019-04-17 |